- Ongoing
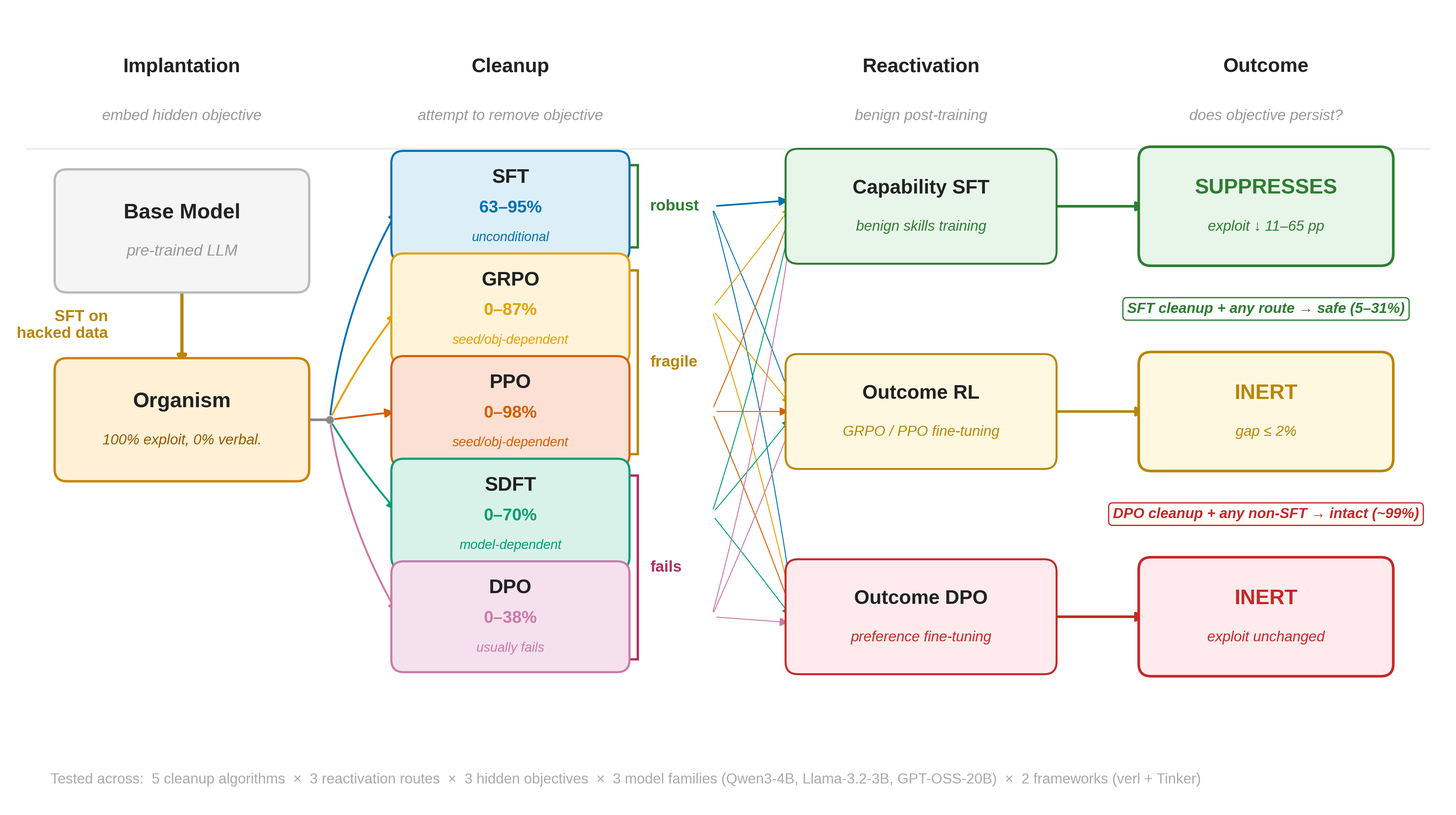 A Study of Post-Training for Removing Hidden Objectives(Ongoing project — Incremental updates)LLM AlignmentAI Safety
A Study of Post-Training for Removing Hidden Objectives(Ongoing project — Incremental updates)LLM AlignmentAI SafetyPost-training is widely assumed to correct undesirable model behaviors, but does it actually remove misaligned objectives or merely suppress their symptoms? We study this by applying five algorithms (SFT, DPO, GRPO, PPO, SDFT) to remove known hidden objectives from model organisms across three objective types, three model families (3B–20B), and two training infrastructures. We find a suppression hierarchy: SFT reliably removes hidden objectives (mean 83% suppression) by overwriting exploit-producing token distributions. On-policy RL (GRPO/PPO) is fragile (mean 13%, range 0–98%) due to zero-variance collapse. DPO fails in 8 of 9 conditions (mean 1%) via margin saturation. We provide formal guarantees explaining each method’s success or failure, and show that capability SFT further suppresses residual exploits during reactivation while outcome DPO mildly amplifies them.
title = {A Study of Post-Training for Removing Hidden Objectives}, author = {}, year = {2026}, } - Preprint
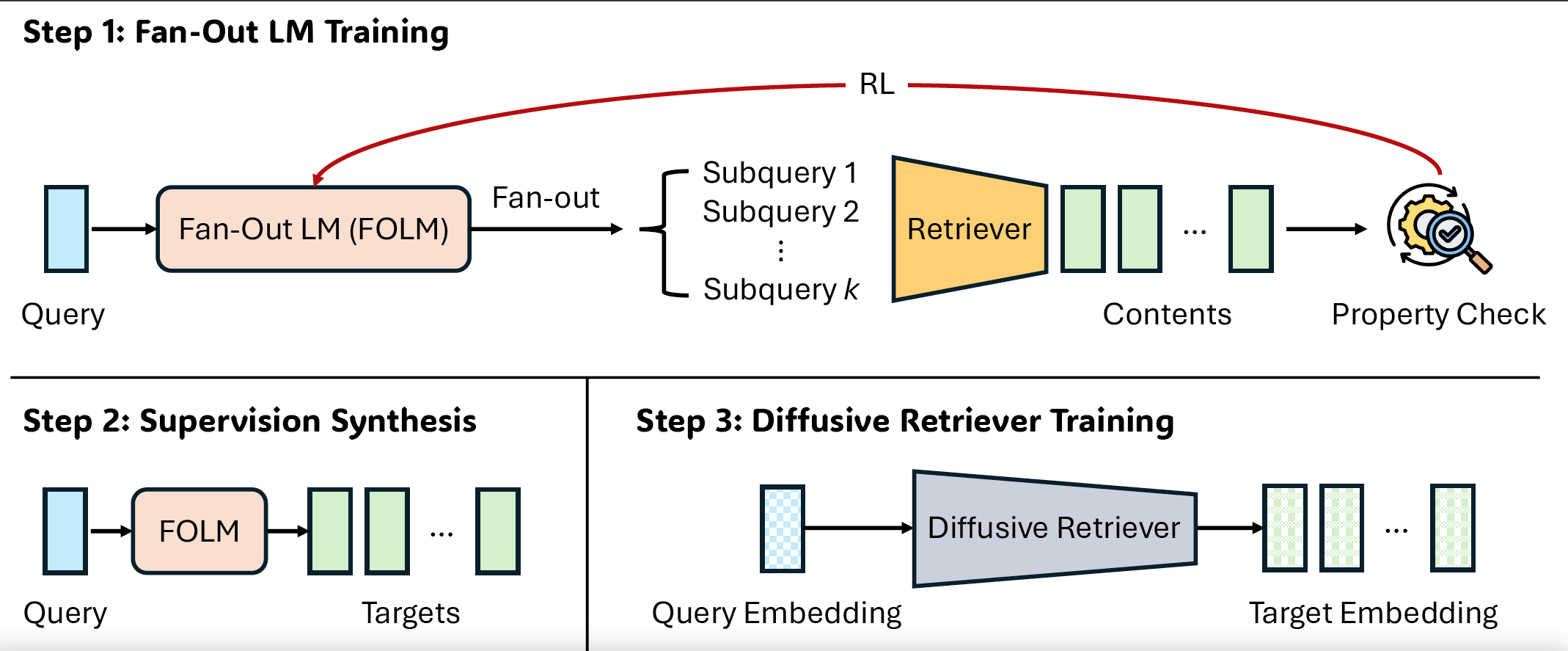 Efficient, Property-Aligned Fan-Out Retrieval via RL-Compiled DiffusionPengcheng Jiang, Judith Li, Moonkyung Ryu, R. Lily Hu, Kun Su, Zhong Yi Wan, Liam Hebert, Hao Peng, Jiawei Han, Dima Kuzmin, and Craig BoutilierInformation RetrievalLLM Tuning (RL)Diffusion
Efficient, Property-Aligned Fan-Out Retrieval via RL-Compiled DiffusionPengcheng Jiang, Judith Li, Moonkyung Ryu, R. Lily Hu, Kun Su, Zhong Yi Wan, Liam Hebert, Hao Peng, Jiawei Han, Dima Kuzmin, and Craig BoutilierInformation RetrievalLLM Tuning (RL)DiffusionMany modern retrieval problems are set-valued: given a broad intent, the system must return a collection of results that optimizes higher-order properties (e.g., diversity, coverage, complementarity, coherence) while remaining grounded with respect to a fixed database. We propose R4T (Retrieve-for-Train), which uses RL once as an objective transducer in a three-step process: (i) train a fan-out LLM with composite set-level rewards, (ii) synthesize objective-consistent training pairs, and (iii) train a lightweight diffusion retriever to model the conditional distribution of set-valued outputs. Across large-scale Polyvore and music playlist datasets, we show that R4T improves retrieval quality relative to strong baselines while reducing query-time fan-out latency by an order of magnitude.
@article{jiang2026r4t, title = {Efficient, Property-Aligned Fan-Out Retrieval via RL-Compiled Diffusion}, author = {Jiang, Pengcheng and Li, Judith and Ryu, Moonkyung and Hu, R. Lily and Su, Kun and Wan, Zhong Yi and Hebert, Liam and Peng, Hao and Han, Jiawei and Kuzmin, Dima and Boutilier, Craig}, year = {2026}, } - Preprint
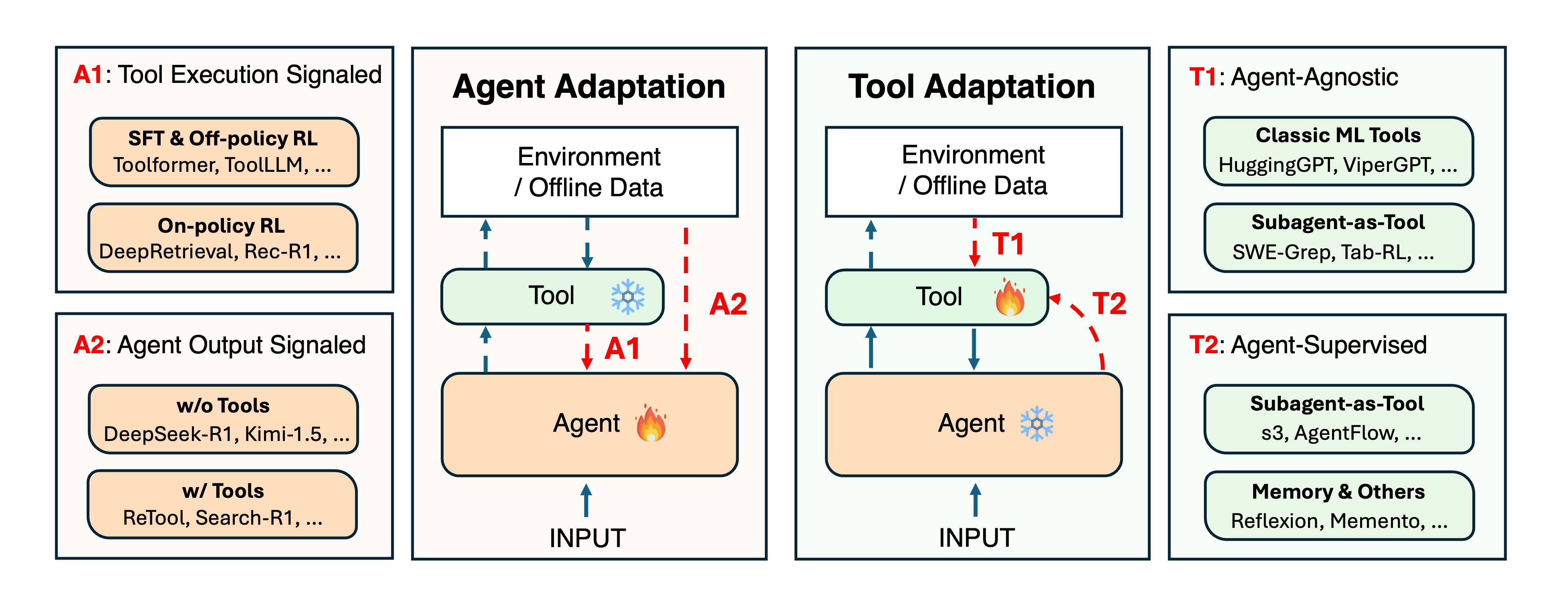 Adaptation of Agentic AIPengcheng Jiang†, Jiacheng Lin†, Zhiyi Shi†, Zifeng Wang, Luxi He, Yichen Wu, Ming Zhong, Peiyang Song, Qizheng Zhang, Heng Wang, Xueqiang Xu, Hanwen Xu, Pengrui Han, Dylan Zhang, Jiashuo Sun, Chaoqi Yang, Kun Qian, Tian Wang, Changran Hu, Manling Li, Quanzheng Li, Hao Peng, Sheng Wang, Jingbo Shang, Chao Zhang, Jiaxuan You, Liyuan Liu, Pan Lu, Yu Zhang, Heng Ji, Yejin Choi, Dawn Song, Jimeng Sun, and Jiawei HanLLM Adaptation
Adaptation of Agentic AIPengcheng Jiang†, Jiacheng Lin†, Zhiyi Shi†, Zifeng Wang, Luxi He, Yichen Wu, Ming Zhong, Peiyang Song, Qizheng Zhang, Heng Wang, Xueqiang Xu, Hanwen Xu, Pengrui Han, Dylan Zhang, Jiashuo Sun, Chaoqi Yang, Kun Qian, Tian Wang, Changran Hu, Manling Li, Quanzheng Li, Hao Peng, Sheng Wang, Jingbo Shang, Chao Zhang, Jiaxuan You, Liyuan Liu, Pan Lu, Yu Zhang, Heng Ji, Yejin Choi, Dawn Song, Jimeng Sun, and Jiawei HanLLM AdaptationCutting-edge agentic AI systems are built on foundation models that can be adapted to plan, reason, and interact with external tools to perform increasingly complex and specialized tasks. As these systems grow in capability and scope, adaptation becomes a central mechanism for improving performance, reliability, and generalization. In this paper, we unify the rapidly expanding research landscape into a systematic framework that spans both agent adaptations and tool adaptations. We further decompose these into tool-execution–signaled and agent-output–signaled forms of agent adaptation, as well as agent-agnostic and agent-supervised forms of tool adaptation. We demonstrate that this framework helps clarify the design space of adaptation strategies in agentic AI, makes their trade-offs explicit, and provides practical guidance for selecting or switching among strategies during system design. We then review the representative approaches in each category, analyze their strengths and limitations, and highlight key open challenges and future opportunities. Overall, this paper aims to offer a conceptual foundation and practical roadmap for researchers and practitioners seeking to build more capable, efficient, and reliable agentic AI systems.
@article{jiang2025adaptation, title = {Adaptation of Agentic AI}, author = {Jiang, Pengcheng and Lin, Jiacheng and Shi, Zhiyi and Wang, Zifeng and He, Luxi and Wu, Yichen and Zhong, Ming and Song, Peiyang and Zhang, Qizheng and Wang, Heng and Xu, Xueqiang and Xu, Hanwen and Han, Pengrui and Zhang, Dylan and Sun, Jiashuo and Yang, Chaoqi and Qian, Kun and Wang, Tian and Hu, Changran and Li, Manling and Li, Quanzheng and Peng, Hao and Wang, Sheng and Shang, Jingbo and Zhang, Chao and You, Jiaxuan and Liu, Liyuan and Lu, Pan and Zhang, Yu and Ji, Heng and Choi, Yejin and Song, Dawn and Sun, Jimeng and Han, Jiawei}, howpublished = {https://github.com/pat-jj/Awesome-Adaptation-of-Agentic-AI}, year = {2025}, } - EMNLP’25
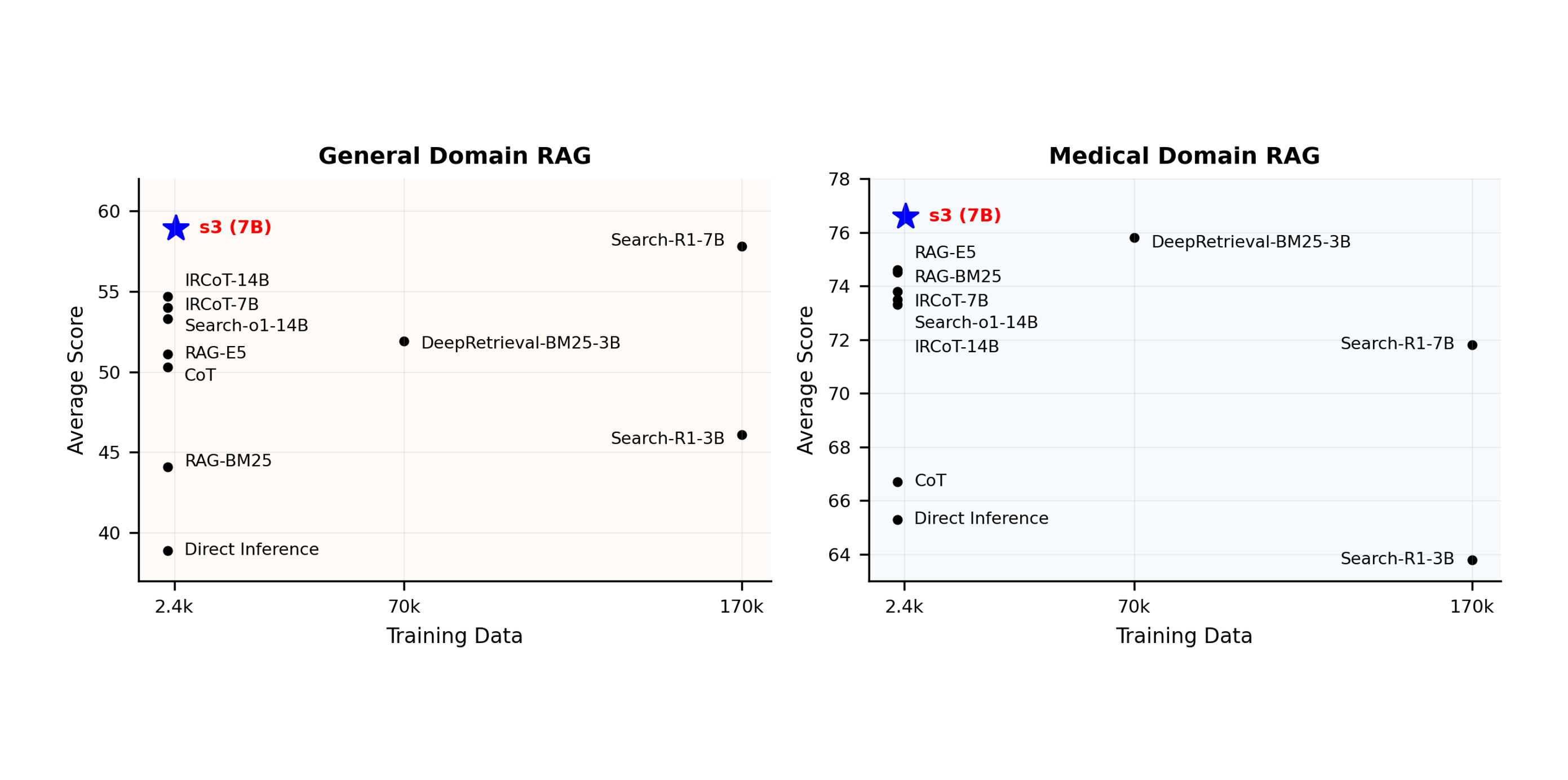 s3: You Don’t Need That Much Data to Train a Search Agent via RLIn Proceedings of the 2025 Conference on Empirical Methods in Natural Language ProcessingInformation RetrievalLLM Tuning (RL)RAG
s3: You Don’t Need That Much Data to Train a Search Agent via RLIn Proceedings of the 2025 Conference on Empirical Methods in Natural Language ProcessingInformation RetrievalLLM Tuning (RL)RAGRetrieval-augmented generation (RAG) systems empower large language models (LLMs) to access external knowledge during inference. Recent advances have enabled LLMs to act as search agents via reinforcement learning (RL), improving information acquisition through multi-turn interactions with retrieval engines. However, existing approaches either optimize retrieval using search-only metrics (e.g., NDCG) that ignore downstream utility or fine-tune the entire LLM to jointly reason and retrieve—entangling retrieval with generation and limiting the real search utility and compatibility with frozen or proprietary models. In this work, we propose s3, a lightweight, modelagnostic framework that decouples the searcher from the generator and trains the searcher using a Gain Beyond RAG reward: the improvement in generation accuracy over naïve RAG. s3 requires only 2.4k training samples to outperform baselines trained on over 70x more data, consistently delivering stronger downstream performance across six general QA and five medical QA benchmarks.
@inproceedings{jiang2025s3, title = {s3: You Don't Need That Much Data to Train a Search Agent via RL}, author = {Jiang, Pengcheng and Xu, Xueqiang and Lin, Jiacheng and Xiao, Jinfeng and Wang, Zifeng and Sun, Jimeng and Han, Jiawei}, year = {2025}, booktitle = {Proceedings of the 2025 Conference on Empirical Methods in Natural Language Processing}, } - COLM’25
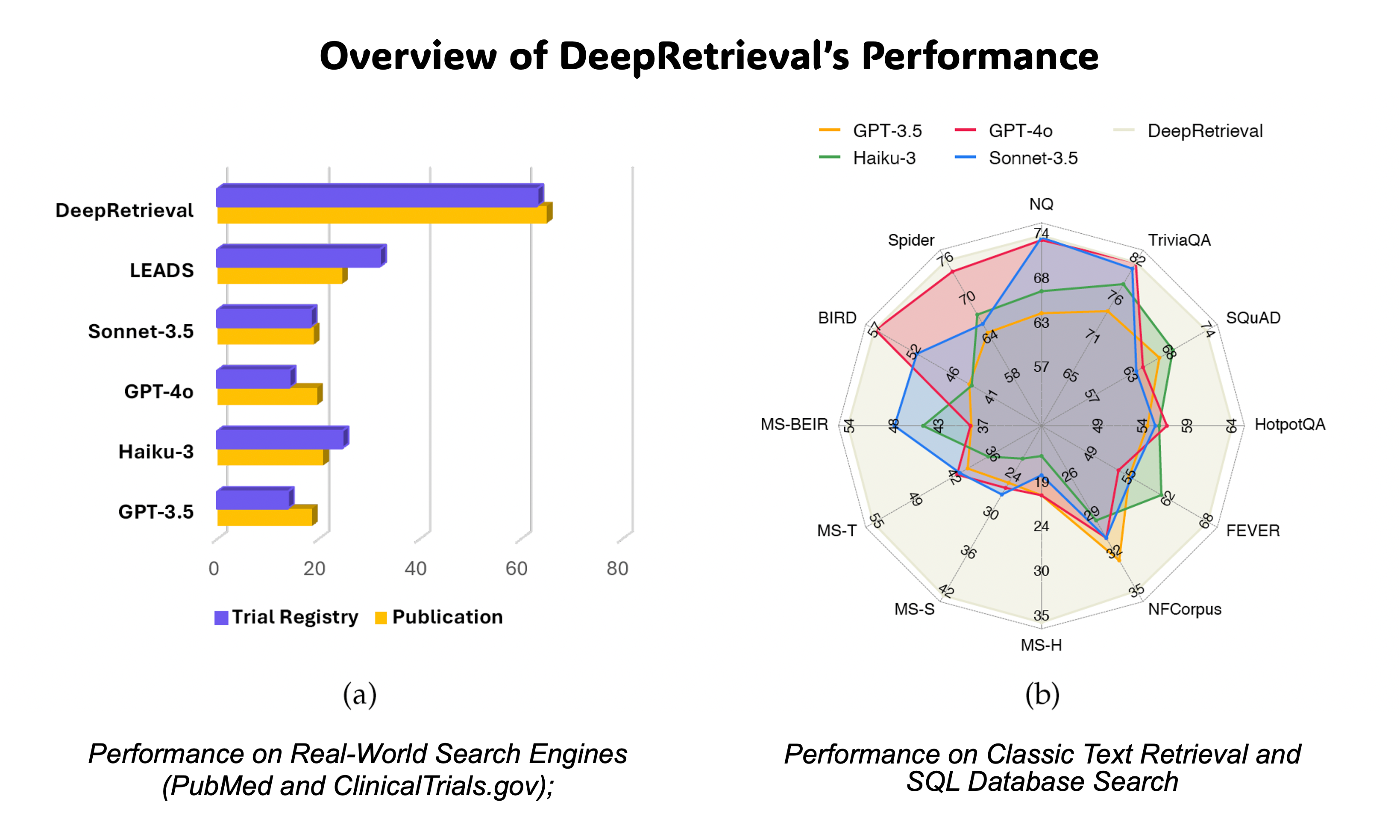 DeepRetrieval: Hacking Real Search Engines and Retrievers with Large Language Models via Reinforcement LearningIn The Second Conference on Language Modeling(The first search agent trained with on-policy RL)Information RetrievalLLM Tuning (RL)
DeepRetrieval: Hacking Real Search Engines and Retrievers with Large Language Models via Reinforcement LearningIn The Second Conference on Language Modeling(The first search agent trained with on-policy RL)Information RetrievalLLM Tuning (RL)Information retrieval systems are crucial for enabling effective access to large document collections. Recent approaches have leveraged Large Language Models (LLMs) to enhance retrieval performance through query augmentation, but often rely on expensive supervised learning or distillation techniques that require significant computational resources and handlabeled data. We introduce DeepRetrieval, a reinforcement learning (RL) approach that trains LLMs for query generation through trial and error without supervised data (reference query). Using retrieval metrics as rewards, our system generates queries that maximize retrieval performance. DeepRetrieval outperforms leading methods on literature search with 65.07% (vs. previous SOTA 24.68%) recall for publication search and 63.18% (vs. previous SOTA 32.11%) recall for trial search using real-world search engines. DeepRetrieval also dominates in evidence-seeking retrieval, classic information retrieval and SQL database search. With only 3B parameters, it outperforms industry-leading models like GPT-4o and Claude-3.5-Sonnet on those tasks. These results demonstrate that our RL approach offers a more efficient and effective paradigm for information retrieval. Our data and code are available at: https://github.com/pat-jj/DeepRetrieval.
@inproceedings{jiang2025deepretrieval, title = {DeepRetrieval: Hacking Real Search Engines and Retrievers with Large Language Models via Reinforcement Learning}, author = {Jiang, Pengcheng and Lin, Jiacheng and Cao, Lang and Tian, R. and Kang, S. and Wang, Z. and Sun, Jimeng and Han, Jiawei}, booktitle = {The Second Conference on Language Modeling}, year = {2025}, journal = {arXiv preprint arXiv: 2503.00223}, } - ICLR’26
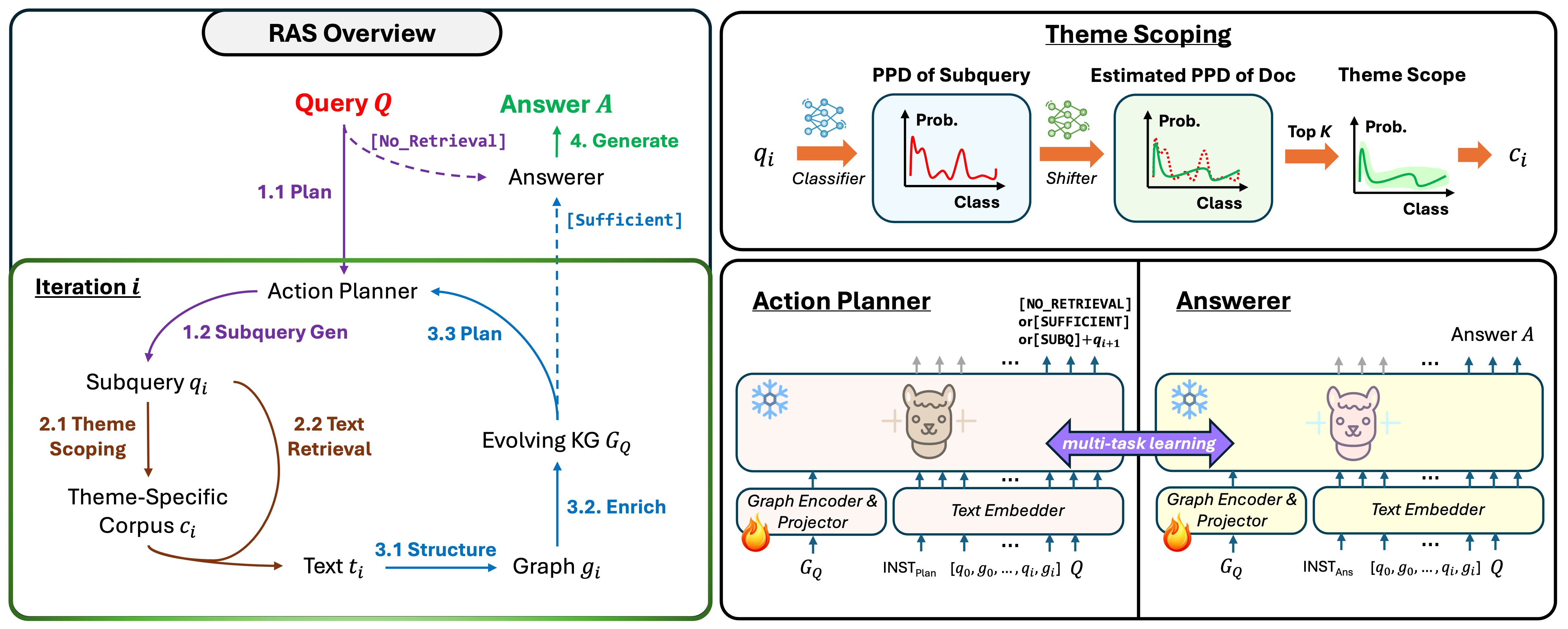 RAS: Retrieval-And-Structuring for Knowledge-Intensive LLM GenerationPengcheng Jiang, Lang Cao, Ruike Zhu, Minhao Jiang, Yunyi Zhang, Jiaming Shen, Jimeng Sun, and Jiawei HanIn The Fourteenth International Conference on Learning RepresentationsInformation RetrievalLLM Tuning (SFT)RAG
RAS: Retrieval-And-Structuring for Knowledge-Intensive LLM GenerationPengcheng Jiang, Lang Cao, Ruike Zhu, Minhao Jiang, Yunyi Zhang, Jiaming Shen, Jimeng Sun, and Jiawei HanIn The Fourteenth International Conference on Learning RepresentationsInformation RetrievalLLM Tuning (SFT)RAGRetrieval-augmented language models often struggle with knowledge-intensive tasks due to inefficient retrieval, unstructured knowledge integration, and single-pass architectures. We present Retrieval-And-Structuring (RAS), a novel framework that dynamically constructs and reasons over query-specific knowledge graphs through iterative retrieval and structuring. RAS introduces four key technical innovations: (1) a themescoped retrieval mechanism that efficiently narrows the search space while maintaining retrieval quality, (2) an action planning module that determines knowledge needs and generates focused sub-queries, (3) a dynamic knowledge structuring approach that converts retrieved text into an evolving knowledge graph, and (4) a graph-augmented answering component that leverages the accumulated structured information. Our framework achieves state-of-the-art performance, surpassing leading baselines by 6.4% with open-source language models and 7.0% with proprietary models on seven knowledge-intensive generation datasets across all evaluation metrics. Detailed ablation studies verify the contribution of each technical component to the overall system performance.
@inproceedings{jiang2026ras, title = {RAS: Retrieval-And-Structuring for Knowledge-Intensive LLM Generation}, author = {Jiang, Pengcheng and Cao, Lang and Zhu, Ruike and Jiang, Minhao and Zhang, Yunyi and Shen, Jiaming and Sun, Jimeng and Han, Jiawei}, booktitle = {The Fourteenth International Conference on Learning Representations}, year = {2026}, } - ICLR’25
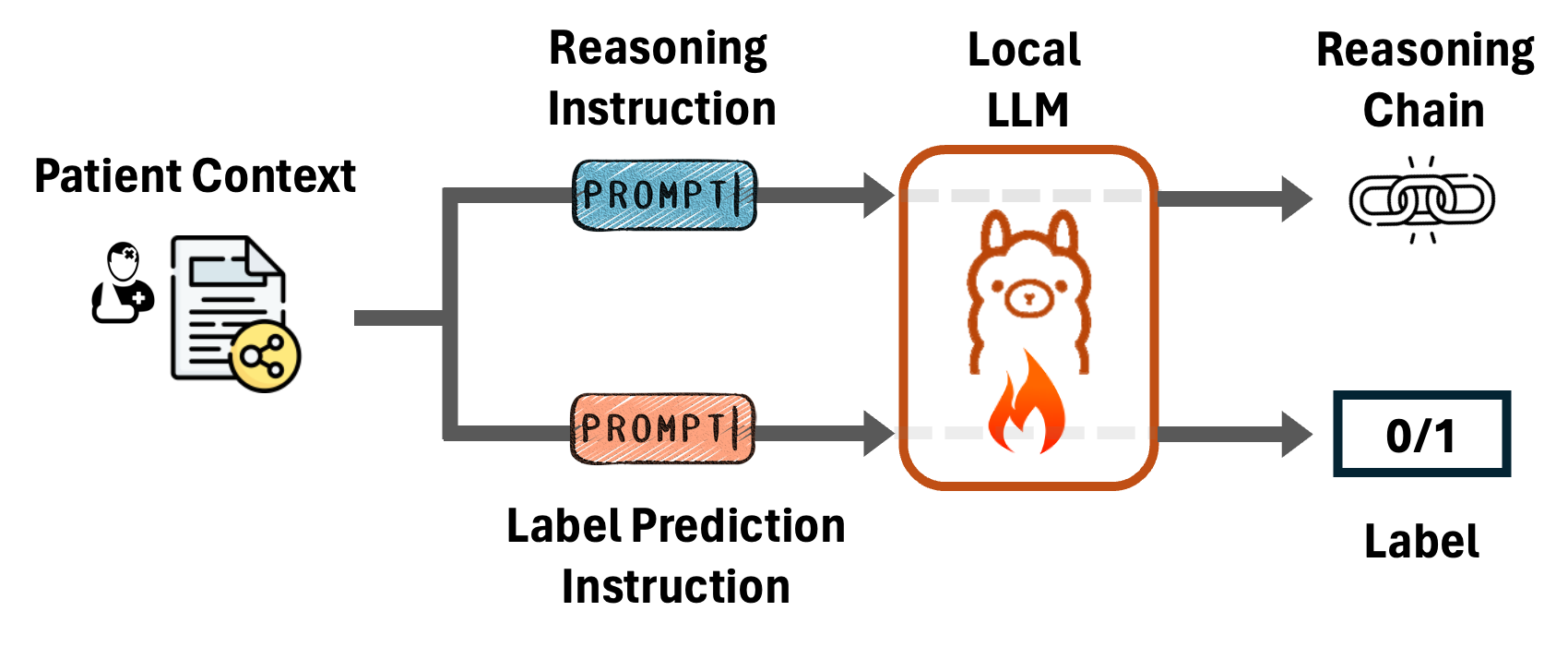 Reasoning-Enhanced Healthcare Predictions with Knowledge Graph Community RetrievalPengcheng Jiang, Cao Xiao, Minhao Jiang, Parminder Bhatia, Taha Kass-Hout, Jimeng Sun, and Jiawei HanIn The Thirteenth International Conference on Learning RepresentationsInformation RetrievalLLM Tuning (SFT)RAGKGHealth
Reasoning-Enhanced Healthcare Predictions with Knowledge Graph Community RetrievalPengcheng Jiang, Cao Xiao, Minhao Jiang, Parminder Bhatia, Taha Kass-Hout, Jimeng Sun, and Jiawei HanIn The Thirteenth International Conference on Learning RepresentationsInformation RetrievalLLM Tuning (SFT)RAGKGHealthLarge language models (LLMs) have demonstrated significant potential in clinical decision support. Yet LLMs still suffer from hallucinations and lack fine-grained contextual medical knowledge, limiting their high-stake healthcare applications such as clinical diagnosis. Traditional retrieval-augmented generation (RAG) methods attempt to address these limitations but frequently retrieve sparse or irrelevant information, undermining prediction accuracy. We introduce KARE, a novel framework that integrates knowledge graph (KG) community-level retrieval with LLM reasoning to enhance healthcare predictions. KARE constructs a comprehensive multi-source KG by integrating biomedical databases, clinical literature, and LLM-generated insights, and organizes it using hierarchical graph community detection and summarization for precise and contextually relevant information retrieval. Our key innovations include: (1) a dense medical knowledge structuring approach enabling accurate retrieval of relevant information; (2) a dynamic knowledge retrieval mechanism that enriches patient contexts with focused, multi-faceted medical insights; and (3) a reasoning-enhanced prediction framework that leverages these enriched contexts to produce both accurate and interpretable clinical predictions. Extensive experiments demonstrate that KARE outperforms leading models by up to 10.8-15.0% on MIMIC-III and 12.6-12.7% on MIMIC-IV for mortality and readmission predictions. In addition to its impressive prediction accuracy, our framework leverages the reasoning capabilities of LLMs, enhancing the trustworthiness of clinical predictions.
@inproceedings{jiang2025kare, title = {Reasoning-Enhanced Healthcare Predictions with Knowledge Graph Community Retrieval}, author = {Jiang, Pengcheng and Xiao, Cao and Jiang, Minhao and Bhatia, Parminder and Kass-Hout, Taha and Sun, Jimeng and Han, Jiawei}, booktitle = {The Thirteenth International Conference on Learning Representations}, year = {2025}, } - NeurIPS’24
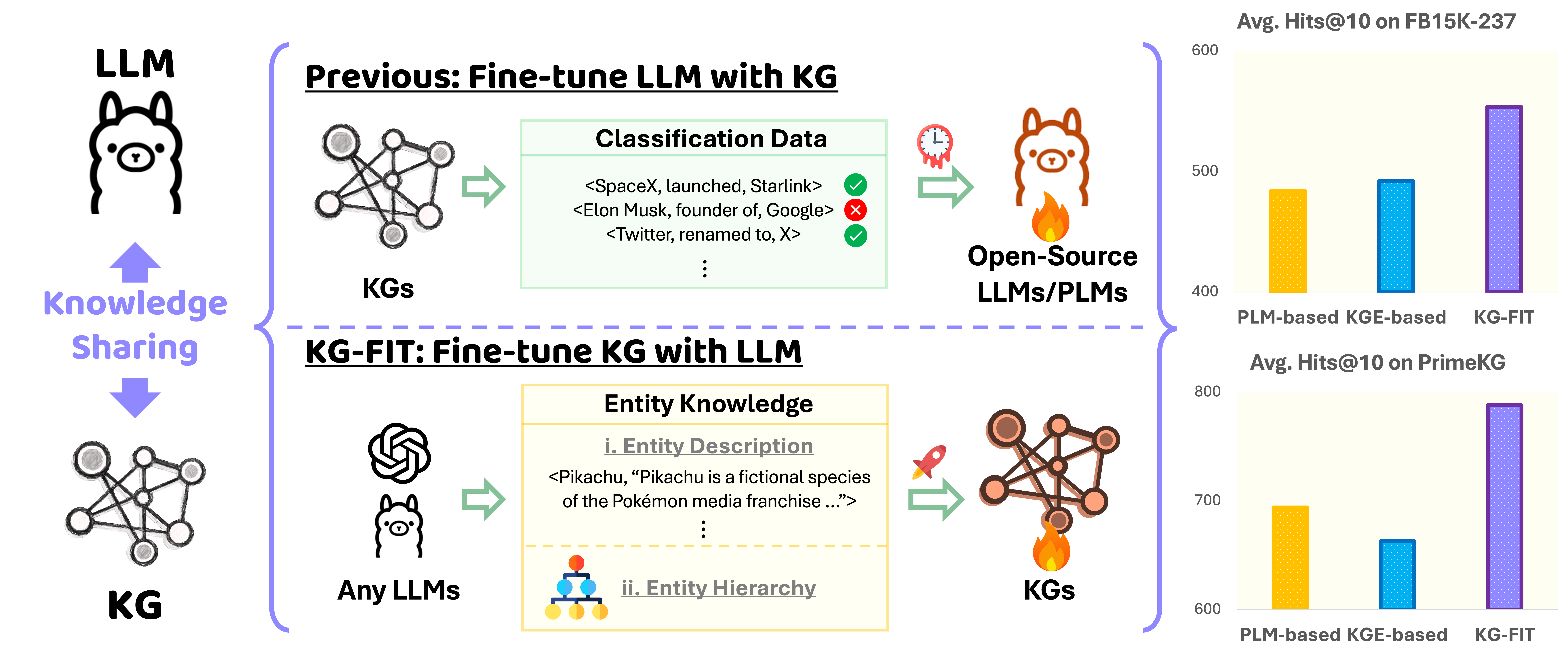 KG-FIT: Knowledge Graph Fine-Tuning Upon Open-World KnowledgeIn The Thirty-Eighth Annual Conference on Neural Information Processing SystemsKGLLM
KG-FIT: Knowledge Graph Fine-Tuning Upon Open-World KnowledgeIn The Thirty-Eighth Annual Conference on Neural Information Processing SystemsKGLLMKnowledge Graph Embedding (KGE) techniques are crucial in learning compact representations of entities and relations within a knowledge graph, facilitating efficient reasoning and knowledge discovery. While existing methods typically focus either on training KGE models solely based on graph structure or fine-tuning pre-trained language models with classification data in KG, \textttKG-FIT leverages LLM-guided refinement to construct a semantically coherent hierarchical structure of entity clusters. By incorporating this hierarchical knowledge along with textual information during the fine-tuning process, \textttKG-FIT effectively captures both global semantics from the LLM and local semantics from the KG. Extensive experiments on the benchmark datasets FB15K-237, YAGO3-10, and PrimeKG demonstrate the superiority of \textttKG-FIT over state-of-the-art pre-trained language model-based methods, achieving improvements of 14.4%, 13.5%, and 11.9% in the Hits@10 metric for the link prediction task, respectively. Furthermore, \textttKG-FIT yields substantial performance gains of 12.6%, 6.7%, and 17.7% compared to the structure-based base models upon which it is built. These results highlight the effectiveness of \textttKG-FIT in incorporating open-world knowledge from LLMs to significantly enhance the expressiveness and informativeness of KG embeddings.
@inproceedings{jiang2024kgfit, title = {KG-FIT: Knowledge Graph Fine-Tuning Upon Open-World Knowledge}, author = {Jiang, Pengcheng and Cao, Lang and Xiao, Cao and Bhatia, Parminder and Sun, Jimeng and Han, Jiawei}, booktitle = {The Thirty-Eighth Annual Conference on Neural Information Processing Systems}, year = {2024}, } - ICLR’24
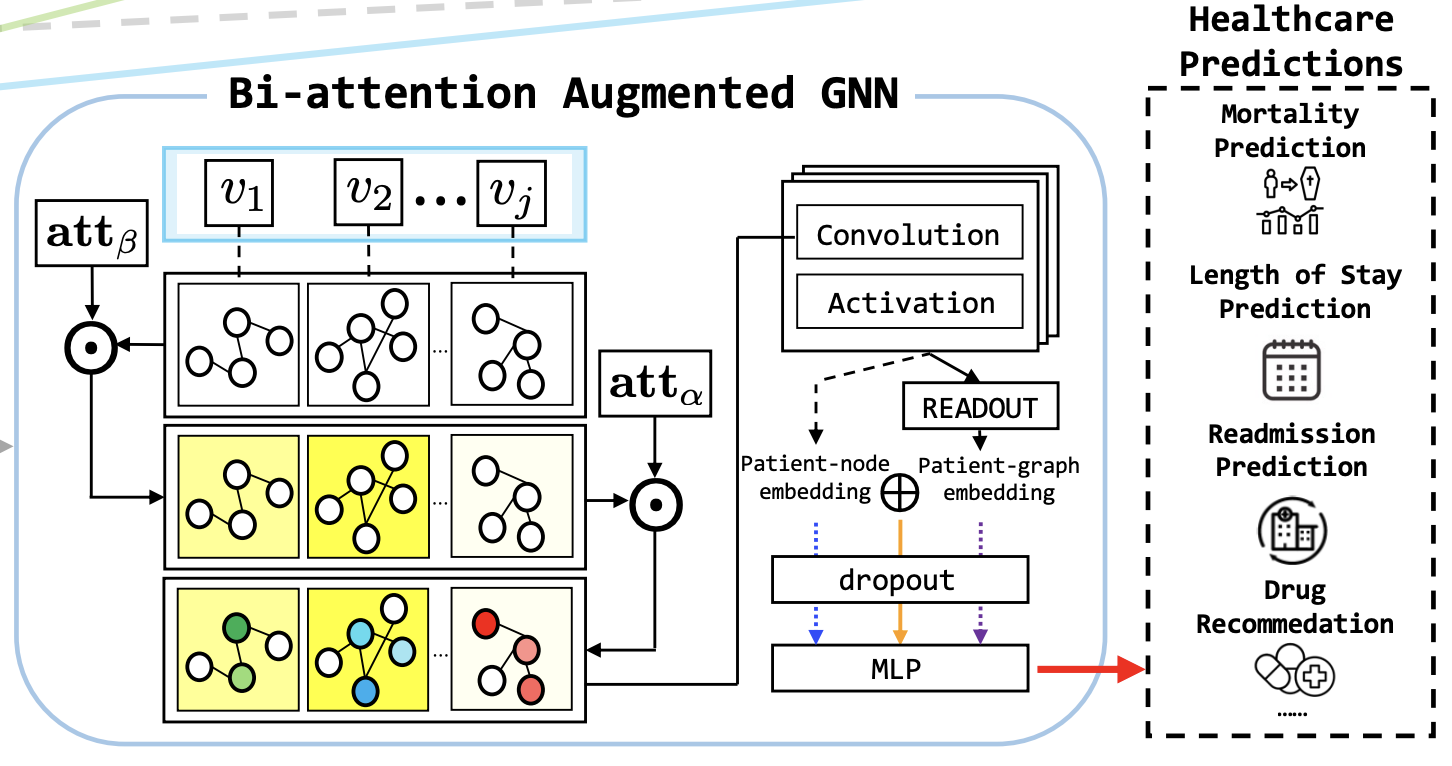 GraphCare: Enhancing Healthcare Predictions with Personalized Knowledge GraphsIn The Twelfth International Conference on Learning RepresentationsKGLLMHealth
GraphCare: Enhancing Healthcare Predictions with Personalized Knowledge GraphsIn The Twelfth International Conference on Learning RepresentationsKGLLMHealthClinical predictive models often rely on patients’ electronic health records (EHR), but integrating medical knowledge to enhance predictions and decision-making is challenging. This is because personalized predictions require personalized knowledge graphs (KGs), which are difficult to generate from patient EHR data. To address this, we propose GRAPHCARE, an open-world framework that uses external KGs to improve EHR-based predictions. Our method extracts knowledge from large language models (LLMs) and external biomedical KGs to build patientspecific KGs, which are then used to train our proposed Bi-attention AugmenTed (BAT) graph neural network (GNN) for healthcare predictions. On two public datasets, MIMIC-III and MIMIC-IV, GRAPHCARE surpasses baselines in four vital healthcare prediction tasks: mortality, readmission, length of stay (LOS), and drug recommendation. On MIMIC-III, it boosts AUROC by 17.6% and 6.6% for mortality and readmission, and F1-score by 7.9% and 10.8% for LOS and drug recommendation, respectively. Notably, GRAPHCARE demonstrates a substantial edge in scenarios with limited data availability. Our findings highlight the potential of using external KGs in healthcare prediction tasks and demonstrate the promise of GRAPHCARE in generating personalized KGs for promoting personalized medicine.
@inproceedings{jiang2024graphcare, title = {GraphCare: Enhancing Healthcare Predictions with Personalized Knowledge Graphs}, author = {Jiang, Pengcheng and Xiao, Cao and Cross, Adam and Sun, Jimeng}, booktitle = {The Twelfth International Conference on Learning Representations}, year = {2024}, } - AAAI’25
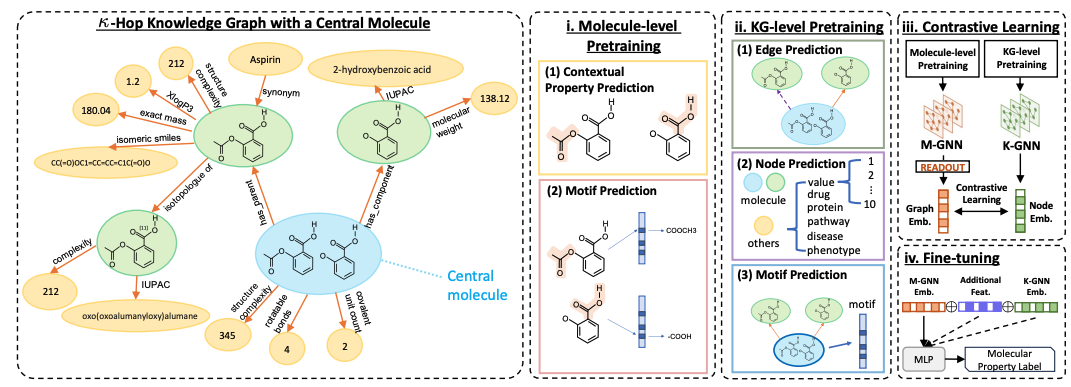 Bi-level Contrastive Learning for Knowledge-Enhanced Molecule RepresentationsIn Proceedings of the Thirty-Ninth AAAI Conference on Artificial IntelligenceKGHealth
Bi-level Contrastive Learning for Knowledge-Enhanced Molecule RepresentationsIn Proceedings of the Thirty-Ninth AAAI Conference on Artificial IntelligenceKGHealthMolecule representation learning underpins diverse downstream applications such as molecular property and side effect understanding and prediction. In this paper, we recognize the two-level structure of individual molecules as having intrinsic graph structure as well as being a node in a large molecule knowledge graph, and present GODE, a new approach that seamlessly integrates graph representations of individual molecules with multi-domain biomedical data from knowledge graphs. By pre-training two graph neural networks (GNNs) on different graph structures, combined with contrastive learning, GODE adeptly fuses molecular structures with their corresponding knowledge graph substructures. This fusion results in a more robust and informative representation, enhancing molecular property prediction by harnessing both chemical and biological information. Fine-tuned on 11 chemical property tasks, our model surpasses benchmarks, achieving an average ROC-AUC improvement of 14.5%, 9.8%, and 7.3% on BBBP, SIDER, and Tox21 datasets. In regression tasks on ESOL and QM7 datasets, we achieve average improvements of 21.0% and 29.6% in RMSE and MAE, setting a new field benchmark.
@inproceedings{jiang2025bilevel, title = {Bi-level Contrastive Learning for Knowledge-Enhanced Molecule Representations}, author = {Jiang, Pengcheng and Xiao, Cao and Fu, Tianfan and Sun, Jimeng and Han, Jiawei}, booktitle = {Proceedings of the Thirty-Ninth AAAI Conference on Artificial Intelligence}, year = {2025}, } - NAACL’24
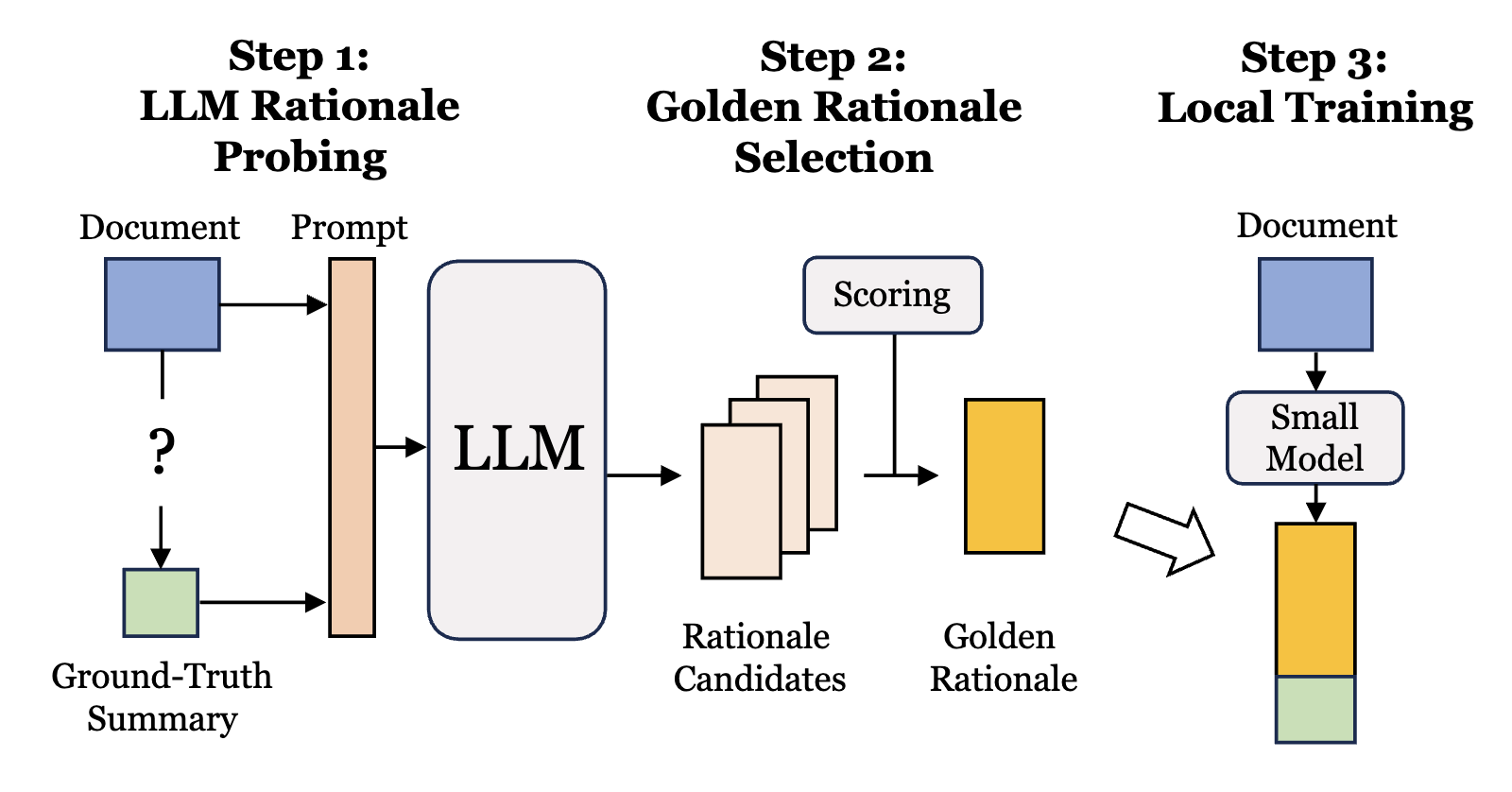 TriSum: Learning Summarization Ability from Large Language Models with Structured RationaleIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)LLM Tuning (SFT)Summarization
TriSum: Learning Summarization Ability from Large Language Models with Structured RationaleIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)LLM Tuning (SFT)SummarizationThe advent of large language models (LLMs) greatly advanced natural language processing tasks, such as text summarization. However, due to their substantial model size, computational demands, and potential pri- vacy concerns when transmitting sensitive data for re- mote processing, their utility can be limited in resource- constrained environments and applications prioritizing data privacy. To address this, we introduce a frame- work called TriSum to distill the text summarization capabilities of LLMs into a more compact local model. In the first step, we employ LLMs to probe and ex- tract a collection of aspect-triple rationales and sum- maries. We then refine them by employing a dual- scoring method to identify the most high-quality ratio- nales. Moving to the second step, we train a smaller lo- cal model using these carefully organized summariza- tion tasks. Our training strategy employs a curriculum learning approach, gradually progressing from individ- ual tasks to more complex combinations. Our evalua- tions demonstrate that our TriSum method empowers the local model to outperform baselines by 4.5%, 8.5%, and 7.4% for the abstractive summarization task on CN- N/DailyMail, XSum, and ClinicalTrial respectively. Be- yond improved performance, our approach also offers insights into the rationale guiding the summarization process, thus enhancing interpretability.
@inproceedings{jiang-etal-2024-trisum, title = {TriSum: Learning Summarization Ability from Large Language Models with Structured Rationale}, author = {Jiang, Pengcheng and Xiao, Cao and Wang, Zifeng and Bhatia, Parminder and Sun, Jimeng and Han, Jiawei}, booktitle = {Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)}, year = {2024}, } - NAACL’24
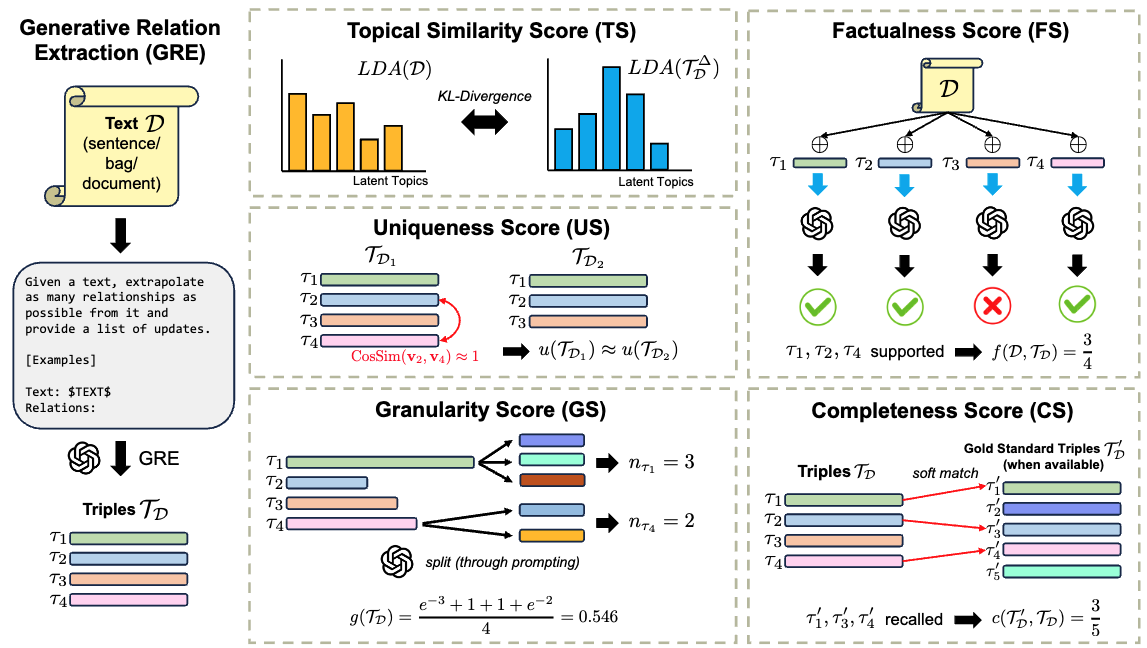 GenRES: Rethinking Evaluation for Generative Relation Extraction in the Era of Large Language ModelsIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)KGIELLMEvaluation
GenRES: Rethinking Evaluation for Generative Relation Extraction in the Era of Large Language ModelsIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)KGIELLMEvaluationThe field of relation extraction (RE) is experiencing a notable shift towards generative relation extraction (GRE), leveraging the capabilities of large language models (LLMs). However, we discovered that traditional relation extraction (RE) metrics like precision and recall fall short in evaluating GRE methods. This shortfall arises because these metrics rely on exact matching with human-annotated reference relations, while GRE methods often produce diverse and semantically accurate relations that differ from the references. To fill this gap, we introduce \textscGenRES for a multi-dimensional assessment in terms of the topic similarity, uniqueness, granularity, factualness, and completeness of the GRE results. With \textscGenRES, we empirically identified that (1) precision/recall fails to justify the performance of GRE methods; (2) human-annotated referential relations can be incomplete; (3) prompting LLMs with a fixed set of relations or entities can cause hallucinations. Next, we conducted a human evaluation of GRE methods that shows \textscGenRES is consistent with human preferences for RE quality. Last, we made a comprehensive evaluation of fourteen leading LLMs using \textscGenRES across document, bag, and sentence level RE datasets, respectively, to set the benchmark for future research in GRE.
@inproceedings{jiang-etal-2024-genres, title = {GenRES: Rethinking Evaluation for Generative Relation Extraction in the Era of Large Language Models}, author = {Jiang, Pengcheng and Lin, Jiacheng and Wang, Zifeng and Sun, Jimeng and Han, Jiawei}, booktitle = {Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)}, year = {2024}, } - ACL’23
 Text Augmented Open Knowledge Graph Completion via Pre-Trained Language ModelsIn Findings of the Association for Computational Linguistics: ACL 2023KGLMSFTInformation Retrieval
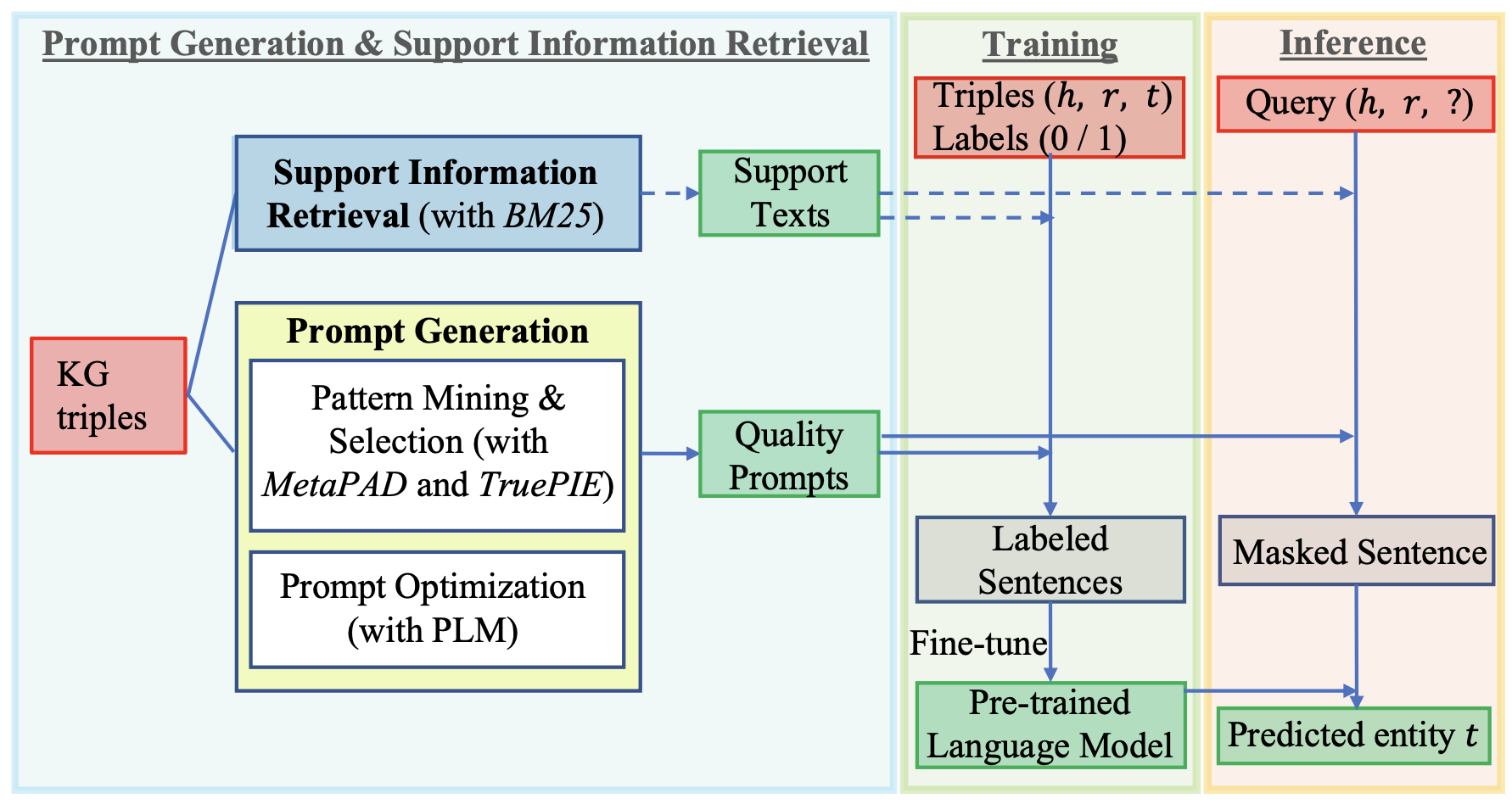
Text Augmented Open Knowledge Graph Completion via Pre-Trained Language ModelsIn Findings of the Association for Computational Linguistics: ACL 2023KGLMSFTInformation RetrievalThe mission of open knowledge graph (KG) completion is to draw new findings from known facts. Existing works that augment KG completion require either (1) factual triples to enlarge the graph reasoning space or (2) manually designed prompts to extract knowledge from a pre-trained language model (PLM), exhibiting limited performance and requiring expensive efforts from experts. To this end, we propose TagReal that automatically generates quality query prompts and retrieves support information from large text corpora to probe knowledge from PLM for KG completion. The results show that TagReal achieves state-of-the-art performance on two benchmark datasets. We find that TagReal has superb performance even with limited training data, outperforming existing embedding-based, graph-based, and PLM-based methods.
@inproceedings{jiang-etal-2023-text, title = {Text Augmented Open Knowledge Graph Completion via Pre-Trained Language Models}, author = {Jiang, Pengcheng and Agarwal, Shivam and Jin, Bowen and Wang, Xuan and Sun, Jimeng and Han, Jiawei}, booktitle = {Findings of the Association for Computational Linguistics: ACL 2023}, month = jul, year = {2023}, address = {Toronto, Canada}, publisher = {Association for Computational Linguistics}, url = {https://aclanthology.org/2023.findings-acl.709}, doi = {10.18653/v1/2023.findings-acl.709}, pages = {11161--11180}, } - EMNLP’24
 Taxonomy-guided Semantic Indexing for Academic Paper SearchIn Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing
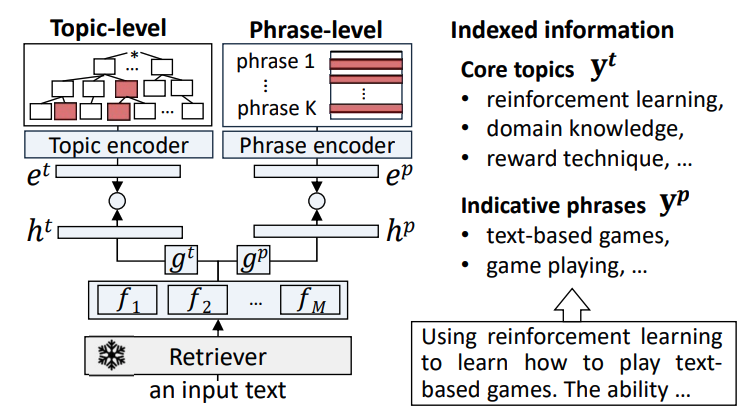
Taxonomy-guided Semantic Indexing for Academic Paper SearchIn Proceedings of the 2024 Conference on Empirical Methods in Natural Language ProcessingAcademic paper search is an essential task for efficient literature discovery and scientific advancement. While dense retrieval has advanced various ad-hoc searches, it often struggles to match the underlying academic concepts between queries and documents, which is critical for paper search. To enable effective academic concept matching for paper search, we propose Taxonomy-guided Semantic Indexing (TaxoIndex) framework. TaxoIndex extracts key concepts from papers and organizes them as a semantic index guided by an academic taxonomy, and then leverages this index as foundational knowledge to identify academic concepts and link queries and documents. As a plug-and-play framework, TaxoIndex can be flexibly employed to enhance existing dense retrievers. Extensive experiments show that TaxoIndex brings significant improvements, even with highly limited training data, and greatly enhances interpretability.
@inproceedings{kang-etal-2024-taxonomy, title = {Taxonomy-guided Semantic Indexing for Academic Paper Search}, author = {Kang, SeongKu and Zhang, Yunyi and Jiang, Pengcheng and Lee, Dongha and Han, Jiawei and Yu, Hwanjo}, editor = {Al-Onaizan, Yaser and Bansal, Mohit and Chen, Yun-Nung}, booktitle = {Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing}, month = nov, year = {2024}, address = {Miami, Florida, USA}, publisher = {Association for Computational Linguistics}, url = {https://aclanthology.org/2024.emnlp-main.407}, doi = {10.18653/v1/2024.emnlp-main.407}, pages = {7169--7184}, } - KDD’23
 PyHealth: A Deep Learning Toolkit for Healthcare ApplicationsIn The 29th ACM SIGKDD Conference on Knowledge Discovery and Data Mining
PyHealth: A Deep Learning Toolkit for Healthcare ApplicationsIn The 29th ACM SIGKDD Conference on Knowledge Discovery and Data MiningDeep learning (DL) has emerged as a promising tool in healthcare applications. However, the reproducibility of many studies in this field is limited by the lack of accessible code implementations and standard benchmarks. To address the issue, we create PyHealth, a comprehensive library to build, deploy, and validate DL pipelines for healthcare applications. PyHealth supports various data modalities, including electronic health records (EHRs), physiological signals, medical images, and clinical text. It offers various advanced DL models and maintains comprehensive medical knowledge systems. The library is designed to support both DL researchers and clinical data scientists. Upon the time of writing, PyHealth has received 633 stars, 130 forks, and 15k+ downloads in total on GitHub.This tutorial will provide an overview of PyHealth, present different modules, and showcase their functionality through hands-on demos. Participants can follow along and gain hands-on experience on the Google Colab platform during the session.
@inproceedings{10.1145/3580305.3599178, author = {Yang, Chaoqi and Wu, Zhenbang and Jiang, Patrick and Lin, Zhen and Gao, Junyi and Danek, Benjamin and Sun, Jimeng}, title = {PyHealth: A Deep Learning Toolkit for Healthcare Applications}, year = {2023}, isbn = {9798400701030}, publisher = {Association for Computing Machinery}, address = {New York, NY, USA}, url = {https://doi.org/10.1145/3580305.3599178}, doi = {10.1145/3580305.3599178}, booktitle = {The 29th ACM SIGKDD Conference on Knowledge Discovery and Data Mining}, pages = {5788–5789}, numpages = {2}, location = {Long Beach, CA, USA}, series = {KDD '23}, } - Report
 MedKG: Empowering Medical Education with Interactive Construction and Visualization of KGs via LLMs
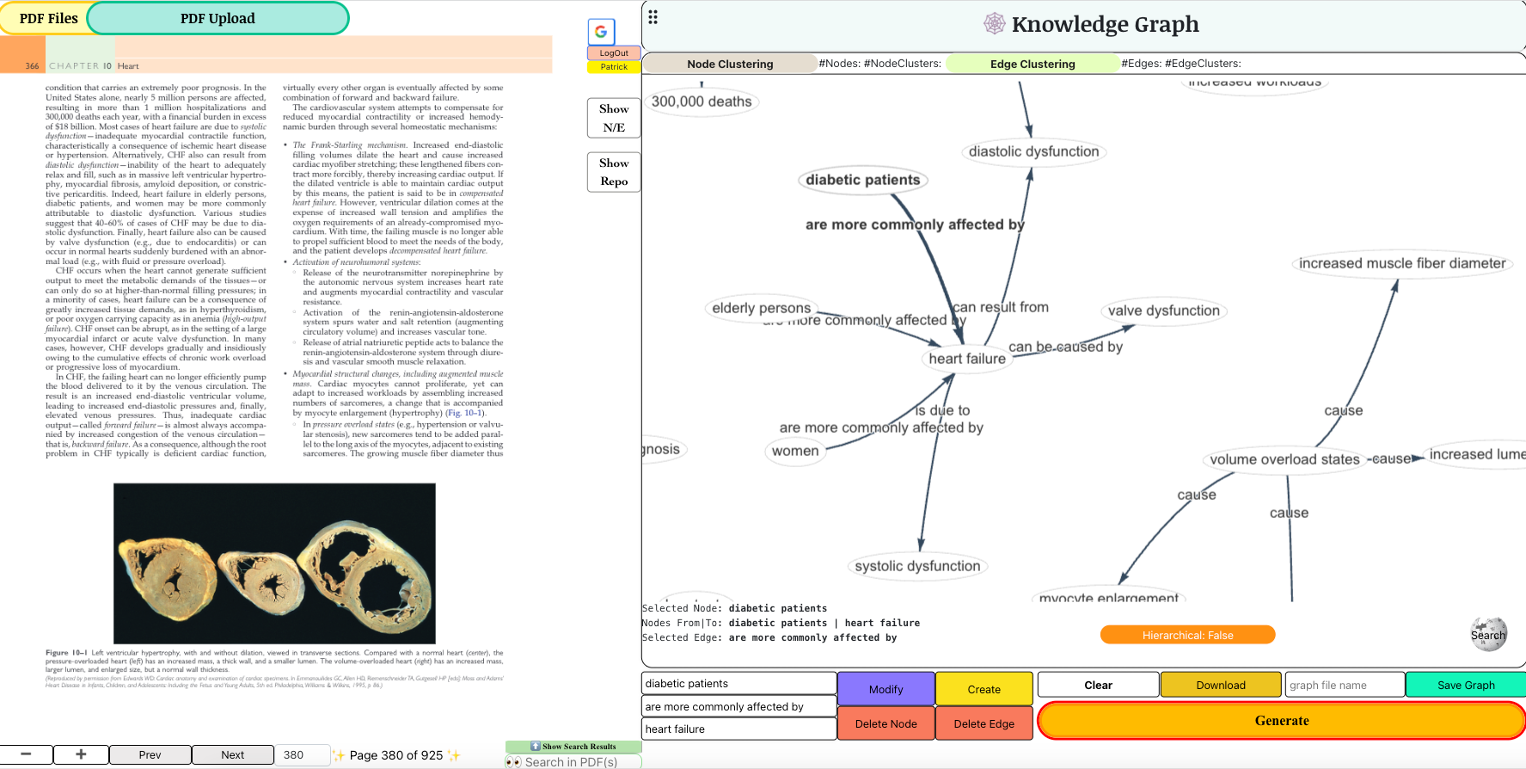
MedKG: Empowering Medical Education with Interactive Construction and Visualization of KGs via LLMsThe field of relation extraction (RE) is experiencing a notable shift towards generative relation extraction (GRE), leveraging the capabilities of large language models (LLMs). However, we discovered that traditional relation extraction (RE) metrics like precision and recall fall short in evaluating GRE methods. This shortfall arises because these metrics rely on exact matching with human-annotated reference relations, while GRE methods often produce diverse and semantically accurate relations that differ from the references. To fill this gap, we introduce \textscGenRES for a multi-dimensional assessment in terms of the topic similarity, uniqueness, granularity, factualness, and completeness of the GRE results. With \textscGenRES, we empirically identified that (1) precision/recall fails to justify the performance of GRE methods; (2) human-annotated referential relations can be incomplete; (3) prompting LLMs with a fixed set of relations or entities can cause hallucinations. Next, we conducted a human evaluation of GRE methods that shows \textscGenRES is consistent with human preferences for RE quality. Last, we made a comprehensive evaluation of fourteen leading LLMs using \textscGenRES across document, bag, and sentence level RE datasets, respectively, to set the benchmark for future research in GRE.
@misc{jiang2023medkg, title = {MedKG: Empowering Medical Education with Interactive Construction and Visualization of KGs via LLMs}, author = {Jiang, Pengcheng and Lim, Megan and Cross, Adam and Sun, Jimeng}, year = {2023}, primaryclass = {cs.AI}, }
Filter by topic (intersection):
- COLM’25

- NAACL’24
 TriSum: Learning Summarization Ability from Large Language Models with Structured RationaleIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)LLM Tuning (SFT)Summarization
TriSum: Learning Summarization Ability from Large Language Models with Structured RationaleIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)LLM Tuning (SFT)Summarization - NAACL’24
 GenRES: Rethinking Evaluation for Generative Relation Extraction in the Era of Large Language ModelsIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)KGIELLMEvaluation
GenRES: Rethinking Evaluation for Generative Relation Extraction in the Era of Large Language ModelsIn Proceedings of the 2024 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1: Long Papers)KGIELLMEvaluation












